Health communication
CONTENTS
Drug-resistant gram-positive bacterial infections
1. Introduction
Antimicrobial resistance (AMR) is a serious threat to global health and occurs when bacteria adapt to withstand the effects of antimicrobial agents. As a result, conventional treatments become ineffective, leading to increased morbidity, mortality, and economic burden.
The misuse and overuse of antibiotics in human medicine, veterinary practice, and agriculture are the major drivers of AMR, including:
- - Use of antibiotics when they are not clinically indicated;
- - Poor-quality medications;
- - Self-medication or inappropriate prescription of antibiotics for viral illnesses;
- - Overuse of antibiotics in hospitals;
- - Use of antibiotics in livestock, facilitating transmission of resistant bacteria from animals to humans;
- - Release of antibiotics into the environment, promoting bacterial evolution and resistance-associated mutations.
According to available estimates:
- Each year, there are 7.7 million deaths associated with bacterial infections, of which 4.95 million are associated with drug-resistant bacteria and 1.27 million are directly attributable to antimicrobial resistance.
- In Europe, the economic burden of AMR is estimated at no less than EUR 1.5 billion annually. In the United States, the total cost reaches USD 55 billion per year.
In 2024, the World Health Organization (WHO) published its global priority list of antimicrobial-resistant pathogens, classified into critical, high, and medium priority groups to guide research and drug development. Among Gram-positive bacteria, the most notable pathogens include:
- Methicillin-resistant Staphylococcus aureus (MRSA): resistant to multiple classes of antibiotics;
- Vancomycin-resistant Enterococcus faecium (VRE): resistant to vancomycin and multiple other agents;
- Streptococcus pneumoniae: categorized as a medium-priority pathogen.
Mechanisms of antimicrobial resistance in Gram-positive bacteria
- - Enzymatic inactivation
- - Alteration of drug targets
- - Reduced drug uptake
- - Increased drug efflux
2. Selected Drug-Resistant Gram-Positive Bacteria
2.1. Staphylococcus aureus
Staphylococcus aureus (S. aureus) is a Gram-positive bacterium belonging to the family Micrococcaceae, commonly colonizing the skin and nasal mucosa in humans. It can cause a broad spectrum of infections involving the skin, soft tissues, and internal organs, some of which may be severe and life-threatening. S. aureus expresses numerous surface adhesion proteins and secretes extracellular products that contribute to both community-acquired and healthcare-associated infections, including surgical site infections and device-associated infections. Food poisoning may also occur following ingestion of food contaminated with staphylococcal toxins. Over the past two decades, the incidence of S. aureus infections has increased substantially in both community and hospital settings. MRSA has been included in the WHO high-priority pathogen list since 2017. In Asia, MRSA is widely prevalent, with very high rates reported in many regions.
Methicillin-susceptible S. aureus (MSSA) is usually treated with oxacillin, nafcillin, cefazolin, or cephalothin; however, S. aureus frequently develops methicillin resistance. For MRSA infections, vancomycin remains a mainstay of therapy.
Major resistance mechanisms of S. aureus
- β-lactamase production: hydrolysis of the β-lactam ring, resulting in high-level resistance to cloxacillin, ceftriaxone, and amoxicillin; even amoxicillin-clavulanate may be less effective when the organism produces large amounts of β-lactamase.
- Aminoglycoside modification: enzymes such as APH/AAC/ANT, encoded by common genes including aac(6')-Ie-aph(2''), aph(3')-IIIa, and ant(4')-Ia, inactivate this drug class and are associated with resistance to gentamicin, tobramycin, kanamycin, and related agents.
- Cell wall alteration and vancomycin target modification:
- VanA/Tn1546 mediates replacement of D-Ala-D-Ala with D-Ala-D-Lac, thereby reducing vancomycin affinity.
- Cell wall thickening creates a “vancomycin trapping” effect, reducing inhibition of peptidoglycan synthesis.
- Altered surface charge mediated by MprF through addition of L-lysine/L-alanine to phosphatidylglycerol reduces susceptibility to cationic peptides and to vancomycin, gentamicin, and daptomycin.
- Biofilm formation: a high proportion of isolates form biofilms; most biofilm-forming strains are multidrug-resistant, and many are MRSA.
- Altered penicillin-binding proteins (mecA/PBP2a): acquisition of mecA, encoding PBP2a with low affinity for β-lactams, confers resistance to methicillin, extended-spectrum penicillins, and cephalosporins and increases the MIC of multiple β-lactams.
- Ribosomal target modification:
- Tet(M)/Tet(S) releases tetracycline from the 30S ribosomal subunit, conferring tetracycline resistance, especially to minocycline via Tet(M).
- Mutations in rplV (L22) and rplD (L4) confer resistance to macrolides and streptogramins.
- Alterations in L3/L4 and mutation rrl5 (23S rRNA, T2500A) are associated with linezolid resistance.
- Efflux pumps: more than 30 potential efflux systems have been described:
- MFS family: Tet(K/L/38/42/43/45/63) for tetracyclines; NorA/NorB/NorC for fluoroquinolones; SdrM, MdeA, and LmrS for multidrug resistance; regulated by MgrA.
- SMR family: QacA/B mediating resistance to quaternary ammonium compounds and chlorhexidine, regulated by QacR; QacC/Ebr, QacD/Smr, and SepA.
- MATE family: MepA, regulated by the mepRAB cluster.
- ABC family: SAV1866 as a structural model and AbcA, associated with hydrophobic β-lactams and autolysis, regulated by MgrA.
-
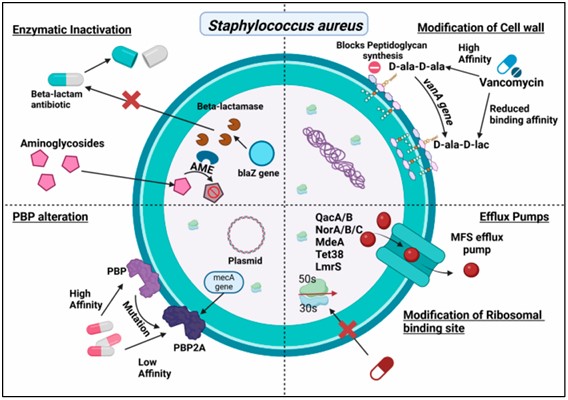
Figure 1. Antibiotic resistance mechanisms of S. aureus
2.2. Streptococcus pneumoniae
Streptococcus pneumoniae (S. pneumoniae) is a Gram-positive bacterium in the family Streptococcaceae and is a major cause of pneumonia, meningitis, otitis media, and other infections. It colonizes the respiratory mucosa and is transmitted by respiratory droplets, facilitating outbreaks. Children, older adults, and immunocompromised individuals are particularly vulnerable. Pneumococci have adapted to multiple antibiotic classes and have progressively developed antimicrobial resistance.
Susceptible strains are treated with penicillin; however, resistance to β-lactams, together with resistance to macrolides, fluoroquinolones, and cotrimoxazole, has become increasingly complex. Pneumococcal meningitis often requires vancomycin-based combination therapy.
Two vaccine types are available: PPV (polysaccharide vaccine) and PCV (conjugate vaccine). Following implementation of PCV7 vaccination in the United States, the incidence of infections caused by penicillin-resistant and multidrug-resistant strains declined substantially, from 6.3 to 2.7 per 100,000 population between 1999 and 2004. Nevertheless, respiratory infections caused by S. pneumoniae remain a major global burden, with WHO estimates suggesting approximately 1.6 million deaths annually.
Major resistance mechanisms of S. pneumoniae
Unlike S. aureus and Enterococcus, S. pneumoniae does not produce β-lactamase, and aminoglycoside-modifying enzymes are less common.
- Altered penicillin-binding proteins: S. pneumoniae is naturally transformable and can acquire mosaic PBPs (PBP1A, PBP1B, PBP2A, PBP2B, PBP2X). Mutations and genetic recombination reduce affinity for β-lactams, especially in PBP1A, PBP2X, and PBP2B, which are associated with high-level penicillin resistance. An example is the PBP2X mutation Gln552→Glu.
- Chromosomal mutations causing fluoroquinolone resistance: resistance usually evolves in two steps, first involving parC (topoisomerase IV), leading to low-level resistance, followed by gyrA (DNA gyrase), resulting in high-level resistance.
- Ribosomal binding-site modification:
- For macrolide-lincosamide-streptogramin resistance, the erm gene mediates methylation of 23S rRNA, reducing drug binding and conferring resistance to erythromycin, azithromycin, clarithromycin, and clindamycin.
- For tetracycline resistance, tet(M) and tet(O) protect the 30S ribosomal subunit and prevent tetracycline binding to the A- and P-sites.
- Efflux pumps:
- PmrA/MFS contributes to reduced fluoroquinolone susceptibility.
- MefE mediates macrolide efflux.
- Biofilm formation: numerous genes are involved in biofilm development and dispersal; biofilms create a favorable environment for horizontal gene transfer (HGT) and maintenance of resistant phenotypes.
-
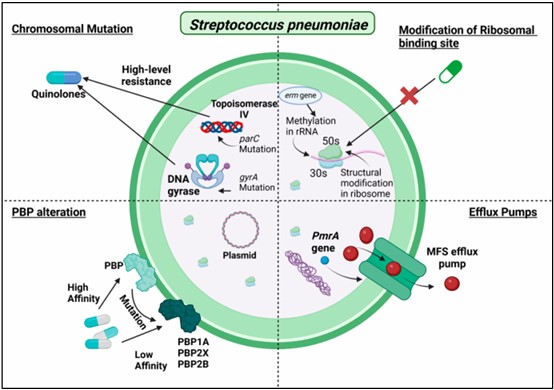
Figure 2. Antibiotic resistance mechanisms of S. pneumoniae
2.3. Enterococcus faecium
Enterococcus faecium (E. faecium) is a Gram-positive coccus that resides in the human gastrointestinal tract and has become an important nosocomial pathogen. It causes urinary tract infections, intra-abdominal infections, endocarditis, and bloodstream infections.
In Europe, Enterococcus ranks second among pathogens causing wound infections and urinary tract infections and third among causes of bloodstream infection. In the United States, approximately 12% of healthcare-associated infections are associated with Enterococcus spp.
This organism has a remarkable capacity for antimicrobial resistance, owing in part to numerous plasmids and transposons carrying resistance genes for erythromycin, gentamicin, kanamycin, streptomycin, tetracycline, and vancomycin. Most vancomycin-resistant E. faecium strains are VRE, with VanA and VanB being the most common phenotypes. Enterococcus also serves as a reservoir of resistance genes that may be transferred to other Gram-positive bacteria, including S. aureus, thereby promoting dissemination of resistance in hospitals.
The organism has also been detected in cheese, fermented milk, beef, pork, poultry, and other foods, suggesting potential foodborne transmission of resistant bacteria to humans.
Resistance mechanisms of E. faecium
- Aminoglycoside modification:
- Genes such as aph(2'')-Ic, aph(2'')-Id, and aph(2'')-Ib confer resistance to amikacin.
- Streptomycin resistance occurs in 50–60% of isolates, and gentamicin resistance in 20–80%.
- Enzymes AAC(6′)-Ii and EfmM confer intrinsic resistance to tobramycin and kanamycin.
- Mutations in ribosomal protein S12 or genes such as ANT(3'')-Ia and ANT(6')-Ia are associated with high-level streptomycin resistance.
- Cell wall target modification: replacement of D-Ala-D-Ala with D-Ala-D-Lac or D-Ala-D-Ser reduces vancomycin affinity.
- VanA: high-level resistance to both vancomycin and teicoplanin (MIC >64 mg/L).
- VanB: variable resistance to vancomycin while retaining susceptibility to teicoplanin.
- VanN: production of D-Ala-D-Ser precursors, conferring intermediate resistance (MIC 8–16 mg/L).
- Biofilm formation: the esp gene is a key determinant facilitating adhesion and persistence in the hospital environment. VRE strains often form robust biofilms and display multidrug resistance to ampicillin, vancomycin, linezolid, and gentamicin.
- Ribosomal target modification:
- Mutation G2576T in 23S rRNA confers linezolid resistance.
- The cfr gene encodes a methyltransferase that methylates A2503 of 23S rRNA, conferring resistance to linezolid, phenicols, lincosamides, and pleuromutilins.
- Mutations in gyrA and parC confer fluoroquinolone resistance.
- Altered penicillin-binding proteins: overexpression or mutation of PBP5 reduces β-lactam affinity, especially for ampicillin.
- Efflux pumps:
- Includes MsrC, Tet(K), and Tet(L) in the MFS family, using proton motive force to export tetracyclines, except tigecycline.
- NorA-like pumps may also contribute to fluoroquinolone resistance.
-
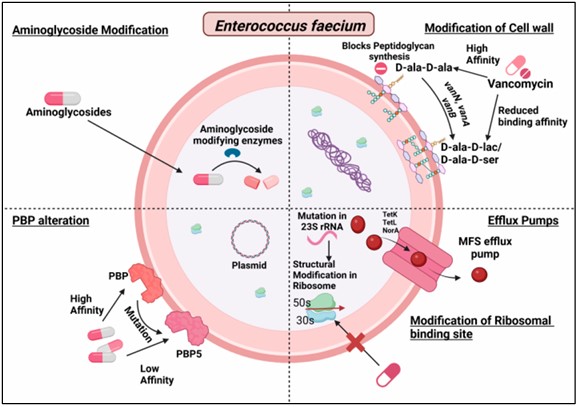
Figure 3. Antibiotic resistance mechanisms of E. faecium
3. Antibiotics for the Treatment of Drug-Resistant Gram-Positive Bacteria
Over the past decade, several new antibiotics have been introduced to address antimicrobial resistance. Newer cephalosporins include ceftobiprole, ceftaroline, cefiderocol, and ceftolozane-tazobactam.
- Ceftobiprole (fifth-generation cephalosporin) is the first β-lactam shown to be effective against MRSA and VRSA because of its strong binding affinity for PBP2a and PBP2x, the latter also being important in S. pneumoniae.
- Delafloxacin (approved in 2017) is a broad-spectrum fluoroquinolone active against MRSA and Pseudomonas aeruginosa and is available in both oral and intravenous formulations for the treatment of acute bacterial skin and skin-structure infections caused by resistant organisms.
- Lefamulin (approved by the FDA in 2019 and by the European Commission in 2020), a pleuromutilin antibiotic, is used to treat community-acquired pneumonia caused by S. pneumoniae and S. aureus; it inhibits protein synthesis by binding to the peptidyl transferase center of the 50S ribosomal subunit.
These newer antibiotics have been developed specifically to overcome existing resistance mechanisms, but they are often reserved as “last-resort” options to reduce the risk of rapid emergence of further resistance. Although some progress has been made, the number of truly effective agents against highly dangerous resistant pathogens remains limited, making real-world assessment difficult and underscoring the urgent need for continued antibiotic research and development.
REFERENCES
Rajput P, Nahar KS, Rahman KM. Evaluation of Antibiotic Resistance Mechanisms in Gram-Positive Bacteria. Antibiotics (Basel). 2024 Dec 8;13(12):1197. doi:10.3390/antibiotics13121197. PMID: 39766587; PMCID: PMC11672434.
MSc. Kim Ngoc Son
MSc. Dinh Thi Thuy Ha

